Golden Gate Assembly¶
What is Golden Gate Assembly?¶
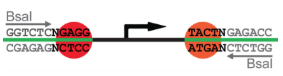
A Golden Gate Assembly is just a restriction / ligation reaction that uses type II restriction enzymes, like BsaI. In these enzymes, the recognition site is different from the cut site, so you can generate any type of overhang for them. See the example below for BsaI, where the recognition site GGTCTC is different from the place they cut.

MoClo / Modular cloning¶
MoClo kits contain plasmids that follow a convention or "syntax". Sets of plasmids within the kit contain the same type of "part". For instance: promoters, coding sequences, RBS, terminators, etc. Plasmids containing parts of the same type, when cut with a type II restriction enzyme, always generate the same overhangs.
For example, in the CIDAR kit, plasmids that contain promoters always produce a fragment containing the promoter flanked with GGAG and ATGA 5' overhangs when cut with BsaI.
Parts of certain types are meant to be assembled together through Golden Gate. The simplest kit would involve the assembly of a promoter, coding sequence, terminator and backbone. Because the overhangs of each part of the assembly are always the same, you can make any combination of parts. In fact, you can create them in a single Golden Gate reaction in what's known as a "Combinatorial Assembly".
Below is the syntax for GoldenPiCS, a combinatorial assembly kit.
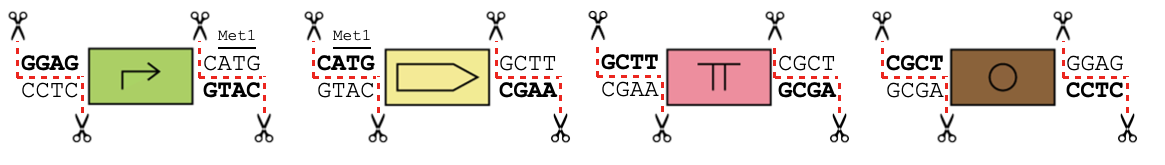
You can load a template to use this kit in OpenCloning here.
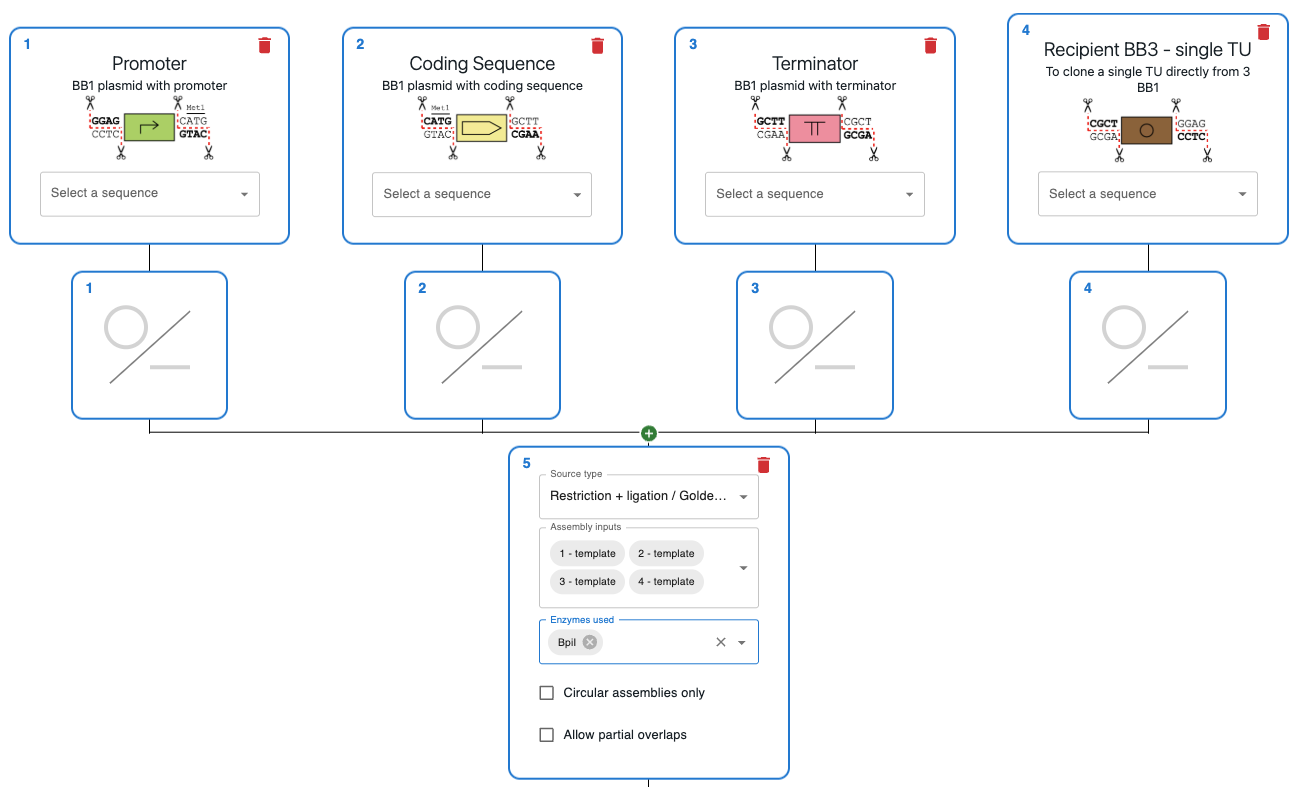
How to plan Golden Gate Assembly using OpenCloning¶
Like any other cloning method, click on the plus icon below a sequence in the Cloning tab and select Restriction + ligation / Golden Gate. Then, select the input sequences in the Assembly inputs field, as well as the enzyme you want to use (you can use more than one). If you know that your desired product is a circular plasmid, tick Circular Assemblies only to exclude other possible linear assemblies from the results.
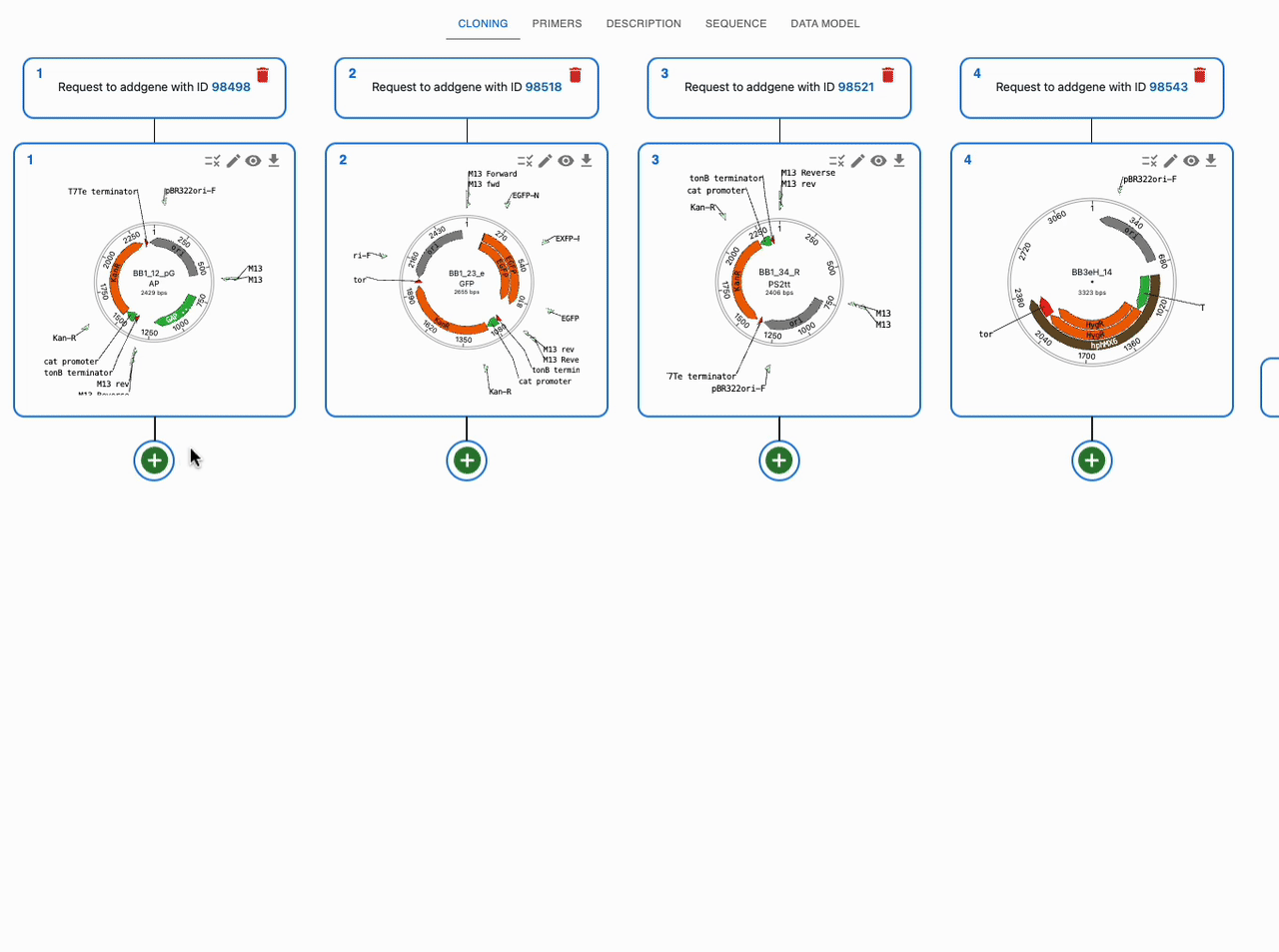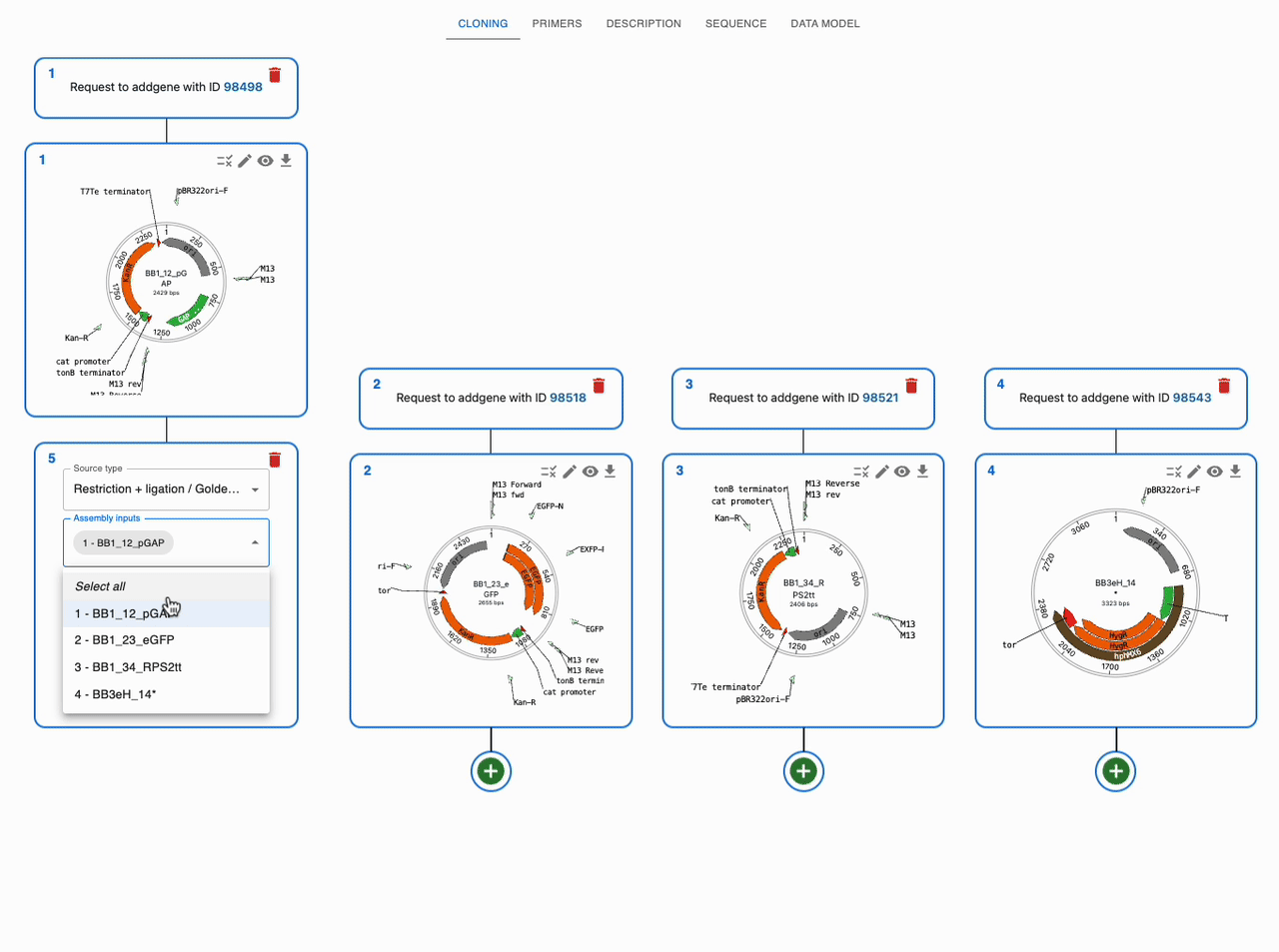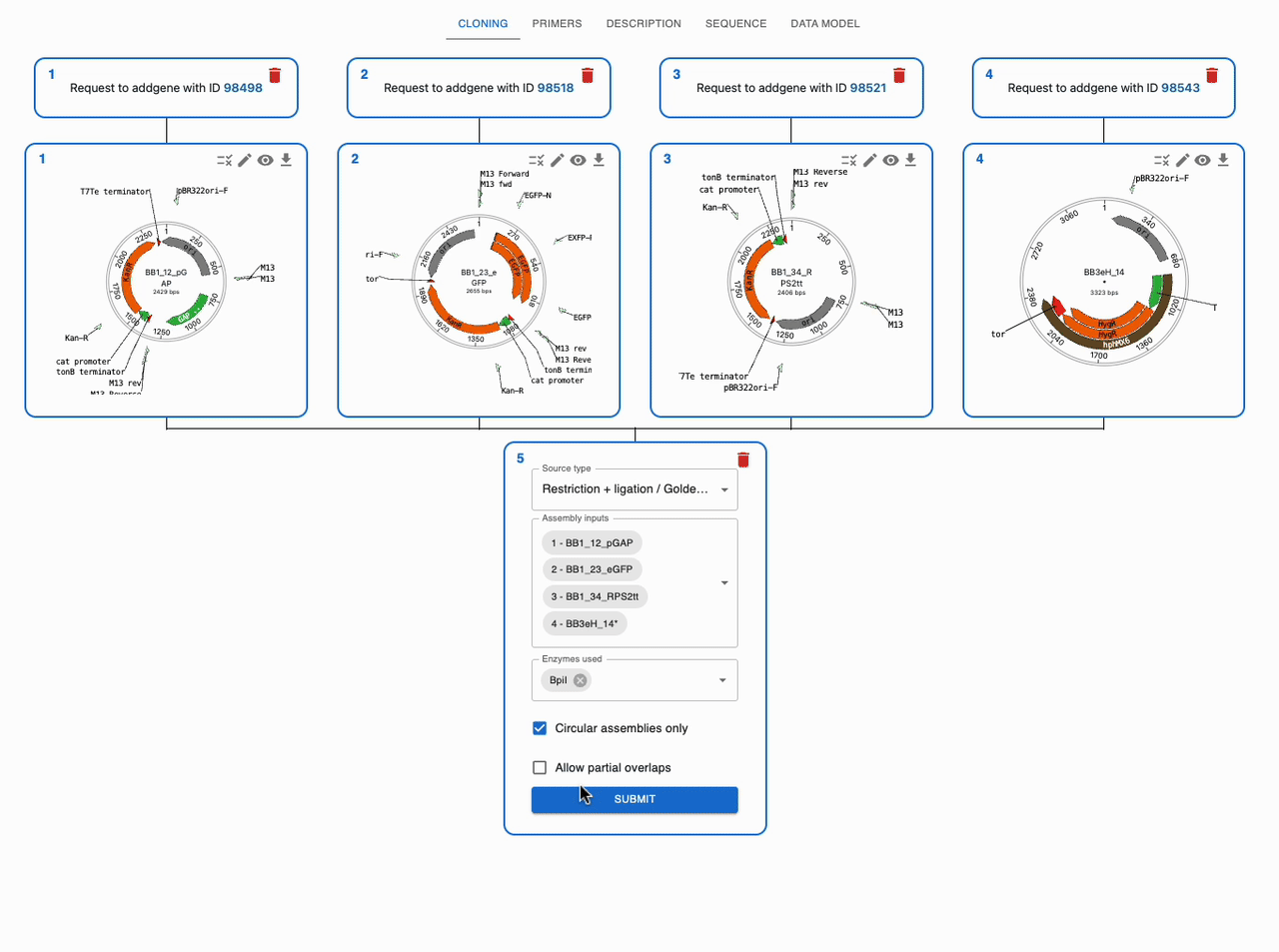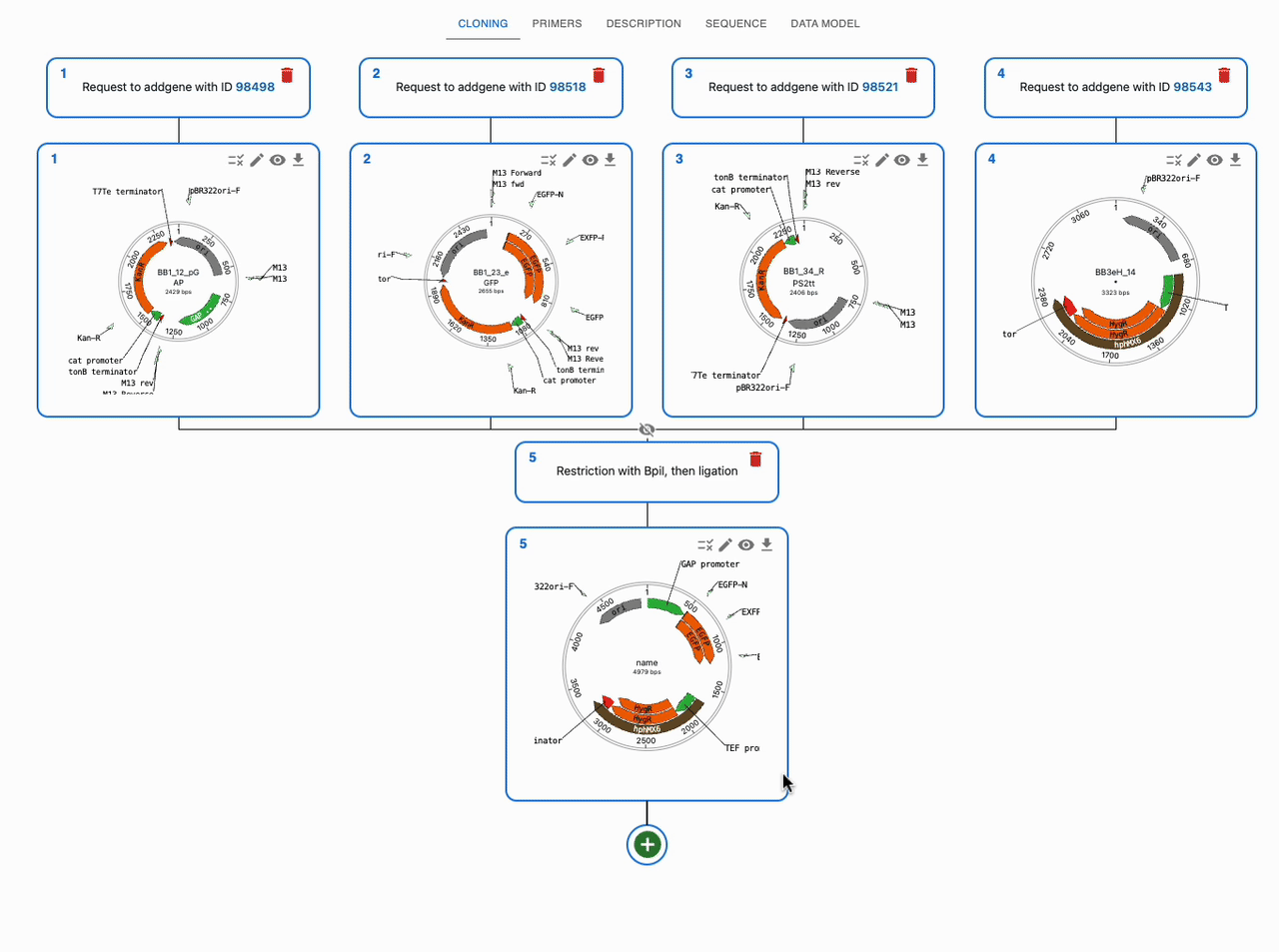
Partially digested products / internal restriction sites¶
If your fragment contains an internal restriction site that you did not notice, the products including it will be filtered out by OpenCloning, because they correspond to partial digestion rather than complete digestion of a single molecule.
If you intentionally want to model the product resulting from partial digestion, you can do so by passing the same input sequence twice. This lets OpenCloning combine products coming from two independently digested copies of the same starting sequence.
Extra features¶
- If you are using a MoClo kit / following a Golden Gate syntax, you can speed up the desing process by using the Assembler.
- If you want to design primers for the Golden Gate Assembly (see Primer design).
- If you want to separate the restriction and ligation steps (see Restriction / Ligation).
- You can pass already digested fragments or Oligonucleotide hybridization products as input sequences. They don't have to be necessarily digested.