Phage Recombinases¶
What is Recombinase-Mediated Cloning?¶
Biological context¶
Recombinase-mediated cloning relies on site-specific recombinases that recognise short DNA sequences and catalyse recombination between them. This is similar to Gateway Cloning and Cre/Lox recombination, but OpenCloning’s Recombinase source lets you define custom recombinases and their recognition sites.
- Recombinases bind to pairs of recognition sites and catalyse recombination between them.
- Many systems use a longer site (attP) and a shorter site (attB), similar to phage integration.

Want to know more?
There is more detailed in the Gateway Cloning page.
How to do Phage Recombinase cloning in OpenCloning?¶
Consensus recognition sites and formatting them¶
Before using your favourite recombinase, you will have to define two consensus recognition sites (equivalent to attP/attB or attL/attR in Gateway Cloning).
- In OpenCloning, they must follow the pattern uppercase–lowercase–uppercase (e.g.
AAaaTTCGGCA,ATGCCCTAAaaCT). - They can include degenerate bases that represent several nucleotides (e.g.
Nfor any nucleotide). - The lowercase letters represent the homology region where sequences are joined, so they must match between the two sites of a pair.
When the two example sites (equivalent to attR and attL) are joined, they will recombine the fragments and generate new sites:
AAaaCT(site1 > site2), likeattB.ATGCCCTAAaaTTCGGCA(site2 > site1), likeattP.
Some recombinases catalyse both the forward and reverse reactions (attB + attP <--> attL + attR). This is possible in OpenCloning by ticking the Reverse reaction option (see below).
Planning the reaction¶
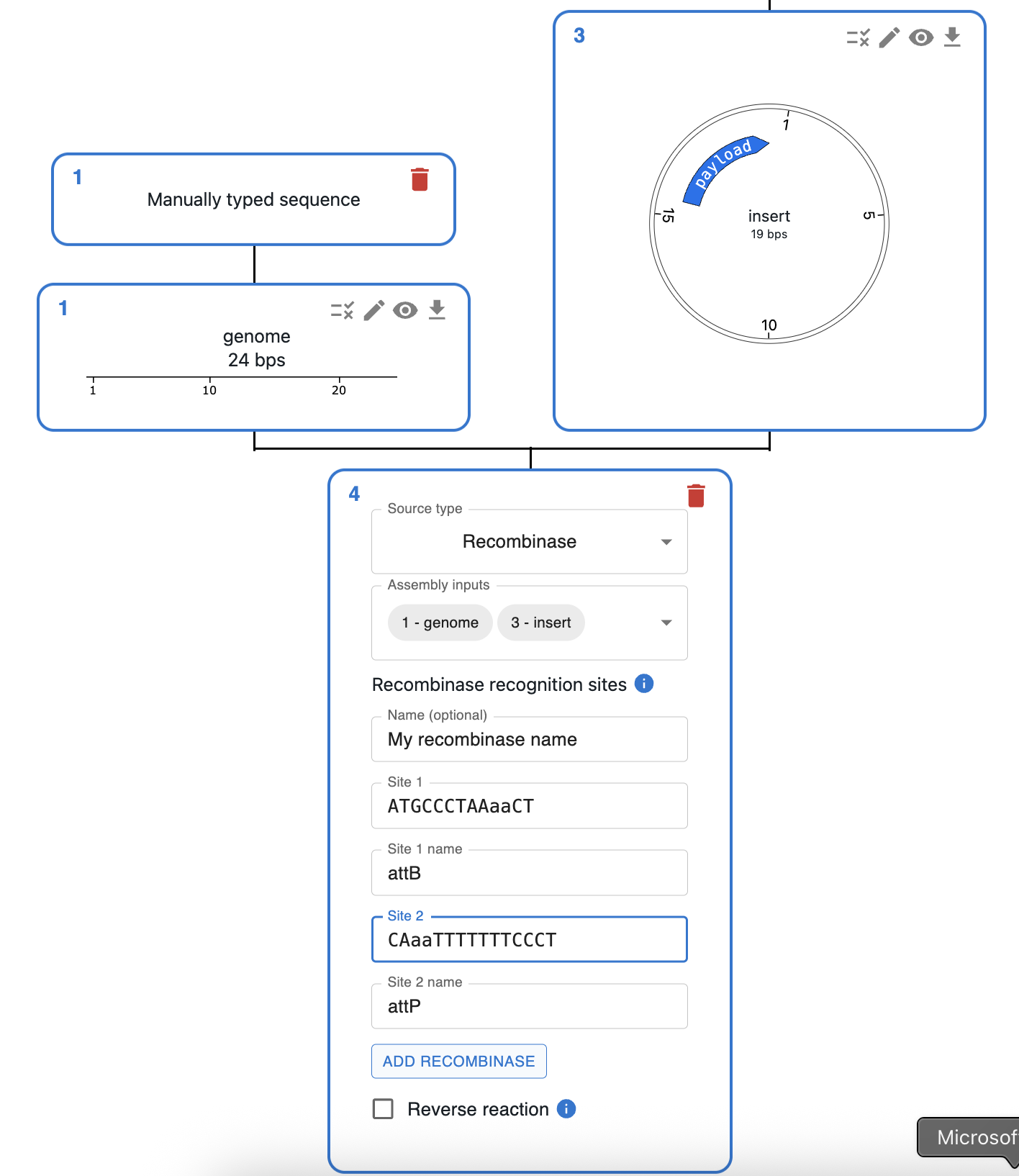
- Like other assembly methods, click the plus icon below a sequence in the
Cloningtab and selectRecombinase. - Select the input sequences in the
Assembly inputsfield. Use one fragment for excision, or two or more for integration. - Add one or more recombinases by specifying their recognition sites. Each recombinase needs:
- Site 1 and Site 2: sequences matching the uppercase–lowercase–uppercase pattern (e.g.
AAaaTTC,CCaaGC). - Site 1 name and Site 2 name (optional): labels such as
attBandattP, which default toattBandattP. - Name (optional): a short name for the recombinase.
- Click on
Add recombinaseto create it. You can use more than one recombinase in the same reaction.
- Site 1 and Site 2: sequences matching the uppercase–lowercase–uppercase pattern (e.g.
- Tick Reverse reaction if you want both forward and reverse recombination to be considered.
- Click Submit to compute the products.